pip install mag45 Komplexeres Plotten & Statistik I [57:38]
In dieser letzten Einheit Daten darzustellen zeige ich – ansatzweise – wie vielfältig mit Jupyter Notebooks, bzw. Python geplottet werden kann, bzw. wie enorm flexibel interaktive Elemente genutzt werden können. Mit dem Package ›geopandas‹ lernen wir außerdem die Möglichkeit kennen, Daten auf Karten darzustellen – natürlich geht auch das interaktiv. Die Bedeutung guter, aussagekräftiger, übersichtlicher, und dabei durchaus komplexer Diagramme kann nicht ausreichend unterstrichen werden. Diagramme und Abbildungen bilden fast immer den Kern eines sehr guten papers, Vortrags oder natürlich einer Abschlussarbeit. Das gelingt jedoch nur mit dem entsprechenden Werkzeug. Matplotlib gehört sicherlich zu den besten Werkzeugen, um wissenschaftliche Diagramme und Abbildungen zu erzeugen. Die hohe Flexibilität interaktiver Elemente erlaubt es sehr schnell und übersichtlich Daten zusammen mit anderen Daten unterschiedlichster Quellen und beliebiger Menge zu visualisieren und analysieren. Das ist ein sehr guter, erster Schritt, um Daten zu verstehen, um dann in die Detail-Analyse zu gehen.
5.1 Program- Populate drop-down menus dynamically [17:01]
.index()
Interaktive Elemente werden dann besonders attraktiv, wenn die darzustellenden Daten, bzw. Dateien ähnlich sind. Dabei genügt es schon, wenn die Kategorienamen in Dateien dieseleben sind, deren Anordnung in den Dateien speilt keine Rolle. Dann wird es möglich, die Inhalte der bspw. Drop-Down Menüs automatisch befüllen zu lassen, um damit einen sehr viel höheren Grad der Flexibilität und dauerhaften Verwendbarkeit des Jupyter Notebooks zu erreichen. Prinzipiell wird es damit auch möglich Notebooks zu schreiben, die man immer verwendet – man passt dann nicht mehr das Notebook an, sondern einfach die darzustellenden Daten. Die Anforderungen dafür sind nicht sonderlich hoch, und es genügt dann, den Datensatz in den entsprechenden Ordner zu kopieren.
| dois | Upload Date | Name | Description | Version | Licence | Keywords | Type | Comments | Short Title | Comment | ORCID | Creation Date | References | Title | Request doi | Source | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | eastsearid | NaN | 0000-0002-5059-2281 | NaN | NaN | Easter Seamount Chain Salas Y Gomez Ridge | NaN | Georoc |
| 1 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | basic | NaN | chondprop | NaN | 0000-0002-5059-2281 | NaN | NaN | Chondrite Properties | NaN | NaN |
| 2 | NaN | 2025-05-16 | Dominik C. Hezel | example dataset | NaN | CCO | chondrules, Fe, isotopes, example data | Example | NaN | chdfeisoexdat | NaN | https://orcid.org/0000-0002-5059-2281 | NaN | NaN | chondrule Fe isotope example data | no | NaN |
| 3 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | basic | NaN | chemelprop | NaN | 0000-0002-5059-2281 | NaN | NaN | Chemical Element Properties | NaN | NaN |
| 4 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | bastcrat | NaN | 0000-0002-5059-2281 | NaN | NaN | Bastar Craton | NaN | Georoc |
| 5 | NaN | 2024-02-06 | Dominik Hezel | Basic data of nuclides | v1.0 | CCO | nuclides, half-lifes, binding-energies | Basic | NaN | nucbasics | NaN | https://orcid.org/0000-0002-5059-2281 | 2024-02-06 | NaN | nuclides-basics | NaN | IAEA - Nuclear Data Section |
| 6 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | ninetyrid | NaN | NaN | NaN | NaN | Ninetyeast Ridge | NaN | Georoc |
| 7 | NaN | 2025-03-14 | Lara Friedrichs | In this table the main elements of the sun's p... | NaN | CC-BY SA | elements, sun, photosphere | Basic | NaN | elements photosphere sun | NaN | https://orcid.org/0009-0001-7264-5081 | 2025-03-14 | NaN | test_table_elements | no | Lodders, K., & Fegley, B. (1998). The planetar... |
| 8 | NaN | 2025-05-19 | Dominik C. Hezel | a brief list of mineral compositions | NaN | CCO | mineral data, example | Basic | NaN | mineraldatex2 | NaN | https://orcid.org/0000-0002-5059-2281 | NaN | NaN | Mineral Data Example Dataset 2 | no | NaN |
| 9 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | emei | NaN | 0000-0002-5059-2281 | NaN | NaN | Emeishan | NaN | Georoc |
| 10 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | mcdis | NaN | 0000-0002-5059-2281 | NaN | NaN | McDonald Islands | NaN | Georoc |
| 11 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | namcord | NaN | 0000-0002-5059-2281 | NaN | NaN | North American Cordillera - Paleozoic | NaN | Georoc |
| 12 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | basic | NaN | etransenergies | NaN | 0000-0002-5059-2281 | NaN | NaN | Element Electron Transition Energies | NaN | NaN |
| 13 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | basic | NaN | chondelab | NaN | 0000-0002-5059-2281 | NaN | NaN | Chondrite Element Abundances | NaN | NaN |
| 14 | NaN | 2025-06-24 | Dominik C. Hezel | movement data for the San Andreas fault in cm | NaN | CCO | creep data | Example | NaN | sanancreep | NaN | https://orcid.org/0000-0002-5059-2281 | NaN | NaN | San Andreas creep | no | NaN |
| 15 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | karaf | NaN | 0000-0002-5059-2281 | NaN | NaN | Karoo Province - Africa | NaN | Georoc |
| 16 | NaN | 2025-05-19 | Dominik C. Hezel | a brief list of mineral compositions | NaN | CCO | mineral data, example | Example | NaN | mineraldatex | NaN | https://orcid.org/0000-0002-5059-2281 | NaN | NaN | Mineral Data Example Dataset | no | NaN |
| 17 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | basic | NaN | ebindenergies | NaN | 0000-0002-5059-2281 | NaN | NaN | Element Electron Binding Energies | NaN | NaN |
| 18 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | galis | NaN | 0000-0002-5059-2281 | NaN | NaN | Galapagos Islands | NaN | Georoc |
| 19 | https://doi.org/10.1180/mgm.2021.43 | 09.12.2023 | Dominik Hezel | IMA–CNMNC approved mineral symbols | NaN | CC-BY-SA | xxx | basic | NaN | abminsym | Another string with\n multiple\n line breaks. | 0000-0002-5059-2281 | 08.06.2021 | Warr LN (2021) Mineralogical Magazine, 85:3, 2... | Abbreviated Mineral Symbols | NaN | paper supplement by L. N. Warr |
| 20 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | hybibplat | NaN | 0000-0002-5059-2281 | NaN | NaN | Hyblean or Iblean Plateau, Sicily | NaN | Georoc |
| 21 | NaN | 2025-05-19 | Dominik C. Hezel | a brief list of mineral compositions | NaN | CCO | mineral data, example | Example | NaN | mineraldatex3 | NaN | https://orcid.org/0000-0002-5059-2281 | NaN | NaN | Mineral Data Example Dataset 3 | no | NaN |
| 22 | NaN | 2024-08-20 | Dominik Hezel | chondrite dataset | NaN | CC-BY SA | chondrite data | Basic | NaN | chondb | NaN | https://orcid.org/0000-0002-5059-2281 | NaN | NaN | chondritedb_test | no | NaN |
| 23 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | basic | NaN | oxelconv | NaN | 0000-0002-5059-2281 | NaN | NaN | Oxide - Element Conversion Factors | NaN | NaN |
| 24 | NaN | 2025-03-13 | Dominik C. Hezel | test | NaN | CCO | test | Basic | NaN | test | NaN | https://orcid.org/0000-0002-5059-2281 | NaN | NaN | mag4_test | no | NaN |
| 25 | NaN | 2024-08-20 | Dominik Hezel | chonddb | NaN | CC-BY SA | chonddb | Basic | NaN | chonddb | NaN | https://orcid.org/0000-0002-5059-2281 | NaN | NaN | chondritedb | no | NaN |
| 26 | NaN | 2025-06-29 | Dominik C. Hezel | Times Series of cps to see variations | NaN | CCO | epma, counts | Example | NaN | epmacounts | NaN | https://orcid.org/0000-0002-5059-2281 | NaN | NaN | EPMA counts for Si Ca Al Ti Fe | no | NaN |
| 27 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | baizone | NaN | 0000-0002-5059-2281 | NaN | NaN | Baical Rift Zone | NaN | Georoc |
| 28 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | westafcrat | NaN | 0000-0002-5059-2281 | NaN | NaN | West African Craton | NaN | Georoc |
| 29 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | bandarc | NaN | 0000-0002-5059-2281 | NaN | NaN | Banda Arc | NaN | Georoc |
| 30 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | tanzcratarch | NaN | 0000-0002-5059-2281 | NaN | NaN | Tanzania Craton Archean | NaN | Georoc |
def pData(xEl, yEl, sel_file1, sel_file2):
df1 = mg.get_data(sel_file1)
df2 = mg.get_data(sel_file2)
plt.scatter(df1[xEl]/10000, df1[yEl]/10000, label=sel_file1)
plt.scatter(df2[xEl]/10000, df2[yEl]/10000, label=sel_file2)
plt.xlabel(xEl + ' (wt%)')
plt.ylabel(yEl + ' (wt%)')
plt.legend()
plt.show()This will always work as long as the category names of all the other databasees are the same – independet from the sorting of the data columns.
✗packagename.dir()
✗packagename.dir
✓dir('packagename')
✗dir = 'packagename'
✓list_name.index(‘element_name_in_list’)
✗list_name.position(‘element_name_in_list’)
✗list_name.element(‘element_name_in_list’)
colon missing in line def
y values are missing
sel_file does not exist as attribute after the command
plt.x must be plt.xlable
plt.x must be plt.ylable
(wt%) needs to be a string
return before plt.show() missing
5.2 Program- Advanced matplotlib plotting [18:30]
fig, ax, subplots(), .subplots_adjust(hspace = 0, wspace = 0), sharex, .set(xlabel = ), .xaxis.set_ticks_position(›both‹), .minorticks_on(), .tick_params(which = ›major‹, length = 7, width = 1, direction = ›in‹), .savefig()
Verwende wie in 5.2 mag4, um Datein zu einzulesen
Jupyter Notebooks, bzw. Python können sehr gut verwendet werden um sehr gute, das bedeutet, sehr übersichtlich, aussagekräftige, und dabei auch komplexe Plots zu erstellen. Es ist praktisch alles möglich – wie meist bei Programmierung –, die Frage ist daher selten ob etwas geht, sondern nur wie das Gewünschte geht. Ich zeige hier ein paar wesentliche Möglichkeiten, welche häufig und typischer für die Mineralogie sind, und verweise sonst auf die tatsächlich diese Woche überarbeitete Dokumentationsseite von https://matplotlib.org, welche mit der Überarbeitung deutlich übersichtlicher und hilfreicher geworden ist (sieht also anders aus als im eben erst erstellten Video). Es lohnt sich schon deshalb, die Seite einmal zu besuchen, um zu sehen, was alles an Diagrammen überhaupt möglich ist. Außerdem zeige ich den Befehl, wie erstellte Abbildungen gespeichert werden können.
| dois | Upload Date | Name | Description | Version | Licence | Keywords | Type | Comments | Short Title | Comment | ORCID | Creation Date | References | Title | Request doi | Source | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | eastsearid | NaN | 0000-0002-5059-2281 | NaN | NaN | Easter Seamount Chain Salas Y Gomez Ridge | NaN | Georoc |
| 1 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | basic | NaN | chondprop | NaN | 0000-0002-5059-2281 | NaN | NaN | Chondrite Properties | NaN | NaN |
| 2 | NaN | 2025-05-16 | Dominik C. Hezel | example dataset | NaN | CCO | chondrules, Fe, isotopes, example data | Example | NaN | chdfeisoexdat | NaN | https://orcid.org/0000-0002-5059-2281 | NaN | NaN | chondrule Fe isotope example data | no | NaN |
| 3 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | basic | NaN | chemelprop | NaN | 0000-0002-5059-2281 | NaN | NaN | Chemical Element Properties | NaN | NaN |
| 4 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | bastcrat | NaN | 0000-0002-5059-2281 | NaN | NaN | Bastar Craton | NaN | Georoc |
| 5 | NaN | 2024-02-06 | Dominik Hezel | Basic data of nuclides | v1.0 | CCO | nuclides, half-lifes, binding-energies | Basic | NaN | nucbasics | NaN | https://orcid.org/0000-0002-5059-2281 | 2024-02-06 | NaN | nuclides-basics | NaN | IAEA - Nuclear Data Section |
| 6 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | ninetyrid | NaN | NaN | NaN | NaN | Ninetyeast Ridge | NaN | Georoc |
| 7 | NaN | 2025-03-14 | Lara Friedrichs | In this table the main elements of the sun's p... | NaN | CC-BY SA | elements, sun, photosphere | Basic | NaN | elements photosphere sun | NaN | https://orcid.org/0009-0001-7264-5081 | 2025-03-14 | NaN | test_table_elements | no | Lodders, K., & Fegley, B. (1998). The planetar... |
| 8 | NaN | 2025-05-19 | Dominik C. Hezel | a brief list of mineral compositions | NaN | CCO | mineral data, example | Basic | NaN | mineraldatex2 | NaN | https://orcid.org/0000-0002-5059-2281 | NaN | NaN | Mineral Data Example Dataset 2 | no | NaN |
| 9 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | emei | NaN | 0000-0002-5059-2281 | NaN | NaN | Emeishan | NaN | Georoc |
| 10 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | mcdis | NaN | 0000-0002-5059-2281 | NaN | NaN | McDonald Islands | NaN | Georoc |
| 11 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | namcord | NaN | 0000-0002-5059-2281 | NaN | NaN | North American Cordillera - Paleozoic | NaN | Georoc |
| 12 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | basic | NaN | etransenergies | NaN | 0000-0002-5059-2281 | NaN | NaN | Element Electron Transition Energies | NaN | NaN |
| 13 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | basic | NaN | chondelab | NaN | 0000-0002-5059-2281 | NaN | NaN | Chondrite Element Abundances | NaN | NaN |
| 14 | NaN | 2025-06-24 | Dominik C. Hezel | movement data for the San Andreas fault in cm | NaN | CCO | creep data | Example | NaN | sanancreep | NaN | https://orcid.org/0000-0002-5059-2281 | NaN | NaN | San Andreas creep | no | NaN |
| 15 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | karaf | NaN | 0000-0002-5059-2281 | NaN | NaN | Karoo Province - Africa | NaN | Georoc |
| 16 | NaN | 2025-05-19 | Dominik C. Hezel | a brief list of mineral compositions | NaN | CCO | mineral data, example | Example | NaN | mineraldatex | NaN | https://orcid.org/0000-0002-5059-2281 | NaN | NaN | Mineral Data Example Dataset | no | NaN |
| 17 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | basic | NaN | ebindenergies | NaN | 0000-0002-5059-2281 | NaN | NaN | Element Electron Binding Energies | NaN | NaN |
| 18 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | galis | NaN | 0000-0002-5059-2281 | NaN | NaN | Galapagos Islands | NaN | Georoc |
| 19 | https://doi.org/10.1180/mgm.2021.43 | 09.12.2023 | Dominik Hezel | IMA–CNMNC approved mineral symbols | NaN | CC-BY-SA | xxx | basic | NaN | abminsym | Another string with\n multiple\n line breaks. | 0000-0002-5059-2281 | 08.06.2021 | Warr LN (2021) Mineralogical Magazine, 85:3, 2... | Abbreviated Mineral Symbols | NaN | paper supplement by L. N. Warr |
| 20 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | hybibplat | NaN | 0000-0002-5059-2281 | NaN | NaN | Hyblean or Iblean Plateau, Sicily | NaN | Georoc |
| 21 | NaN | 2025-05-19 | Dominik C. Hezel | a brief list of mineral compositions | NaN | CCO | mineral data, example | Example | NaN | mineraldatex3 | NaN | https://orcid.org/0000-0002-5059-2281 | NaN | NaN | Mineral Data Example Dataset 3 | no | NaN |
| 22 | NaN | 2024-08-20 | Dominik Hezel | chondrite dataset | NaN | CC-BY SA | chondrite data | Basic | NaN | chondb | NaN | https://orcid.org/0000-0002-5059-2281 | NaN | NaN | chondritedb_test | no | NaN |
| 23 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | basic | NaN | oxelconv | NaN | 0000-0002-5059-2281 | NaN | NaN | Oxide - Element Conversion Factors | NaN | NaN |
| 24 | NaN | 2025-03-13 | Dominik C. Hezel | test | NaN | CCO | test | Basic | NaN | test | NaN | https://orcid.org/0000-0002-5059-2281 | NaN | NaN | mag4_test | no | NaN |
| 25 | NaN | 2024-08-20 | Dominik Hezel | chonddb | NaN | CC-BY SA | chonddb | Basic | NaN | chonddb | NaN | https://orcid.org/0000-0002-5059-2281 | NaN | NaN | chondritedb | no | NaN |
| 26 | NaN | 2025-06-29 | Dominik C. Hezel | Times Series of cps to see variations | NaN | CCO | epma, counts | Example | NaN | epmacounts | NaN | https://orcid.org/0000-0002-5059-2281 | NaN | NaN | EPMA counts for Si Ca Al Ti Fe | no | NaN |
| 27 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | baizone | NaN | 0000-0002-5059-2281 | NaN | NaN | Baical Rift Zone | NaN | Georoc |
| 28 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | westafcrat | NaN | 0000-0002-5059-2281 | NaN | NaN | West African Craton | NaN | Georoc |
| 29 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | bandarc | NaN | 0000-0002-5059-2281 | NaN | NaN | Banda Arc | NaN | Georoc |
| 30 | NaN | 21.01.2024 | Dominik Hezel | xxx | NaN | CC-BY-SA | xxx | Database Dataset | NaN | tanzcratarch | NaN | 0000-0002-5059-2281 | NaN | NaN | Tanzania Craton Archean | NaN | Georoc |
['Easter Seamount Chain Salas Y Gomez Ridge',
'Bastar Craton',
'Ninetyeast Ridge',
'Emeishan',
'McDonald Islands',
'North American Cordillera - Paleozoic',
'Karoo Province - Africa',
'Galapagos Islands',
'Hyblean or Iblean Plateau, Sicily',
'Baical Rift Zone',
'West African Craton',
'Banda Arc',
'Tanzania Craton Archean']df1 = mg.get_data('Bastar Craton')
df2 = mg.get_data('Banda Arc')
df3 = mg.get_data('Ninetyeast Ridge')
df4 = mg.get_data('McDonald Islands')
xEl = 'Al'
yEl = 'Mg'
fac = .0001
fig, ((ax1, ax2), (ax3, ax4)) = plt.subplots(2, 2, sharex = True, sharey = True)
fig.subplots_adjust(hspace = 0, wspace = 0)
ax1.scatter(df1[xEl] * fac, df1[yEl] * fac)
ax2.scatter(df2[xEl] * fac, df2[yEl] * fac)
ax3.scatter(df3[xEl] * fac, df3[yEl] * fac)
ax4.scatter(df4[xEl] * fac, df4[yEl] * fac)
ax1.set(ylabel = yEl + ' wt%')
ax3.set(xlabel = xEl + ' wt%', ylabel = yEl + ' wt%')
ax4.set(xlabel = xEl + ' wt%')
ax1.xaxis.set_ticks_position('both')
ax1.yaxis.set_ticks_position('both')
ax2.xaxis.set_ticks_position('both')
ax2.yaxis.set_ticks_position('both')
ax1.minorticks_on()
ax2.minorticks_on()
ax1.tick_params(which = 'major', length = 7, width = 1, direction = 'in')
ax1.tick_params(which = 'minor', length = 4, width = 1, direction = 'in')
fig.savefig('test.pdf')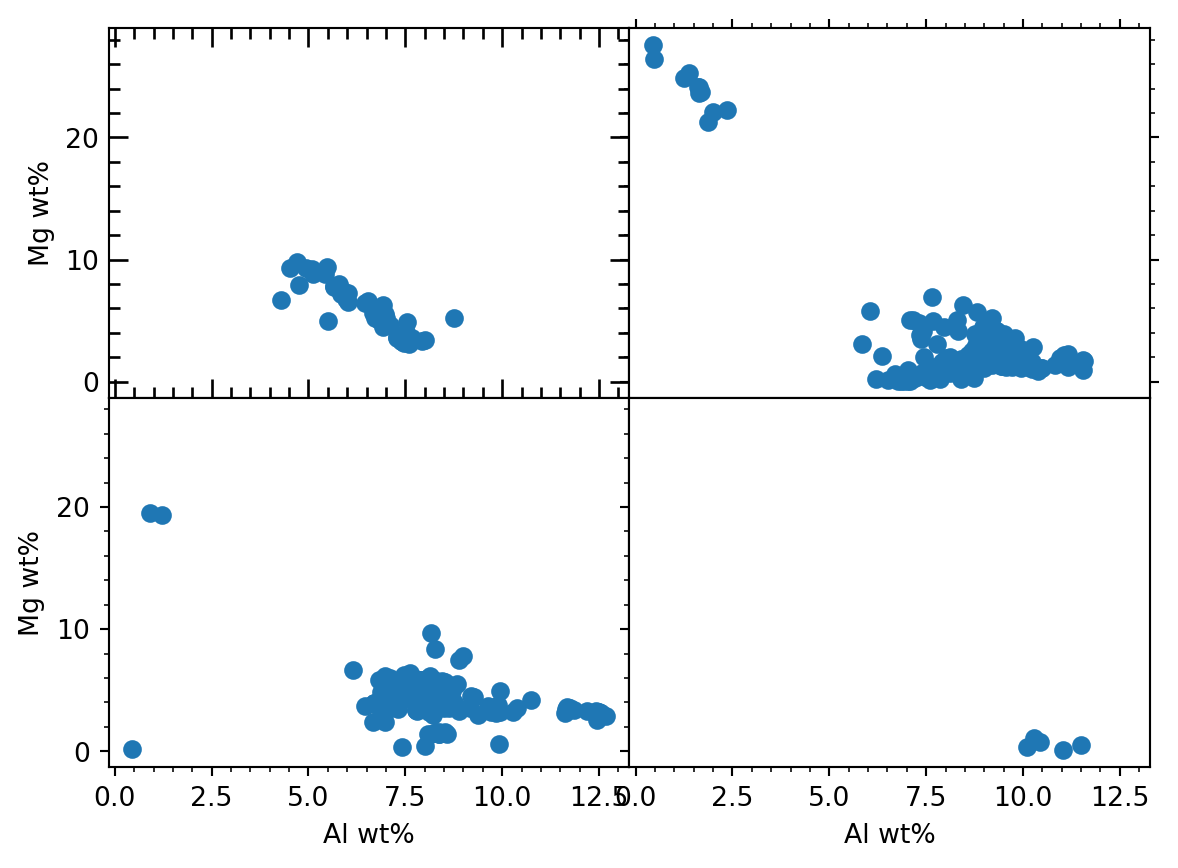
A figure (fig) can contain multiple plots. A plot (plt) represents only a single plot.
✓True
✗False
✗True
✓False
import matplotlib.pyplot as plt
space = True
if space == True:
wsWidth = .1
else:
wsWidth = 0
fig, ((ax1, ax2), (ax3, ax4)) = plt.subplots(2, 2, sharey = True)
fig.subplots_adjust(hspace = 0, wspace = wsWidth)
ax1.scatter([1,2,3], [4,5,6])
ax2.scatter([7,8,9], [1,2,3])
ax3.scatter([4,5,6], [7,8,9])
ax4.scatter([3,2,1], [4,5,6])
plt.show()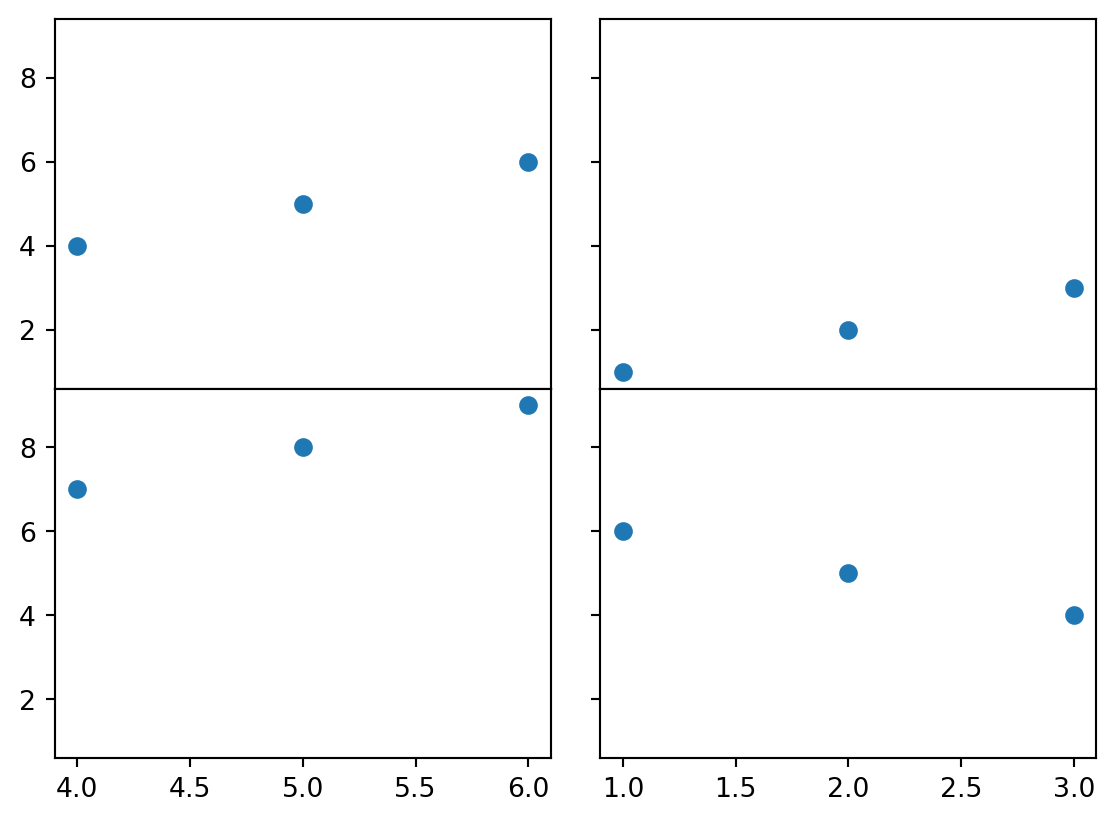
The goal is have 6 plots in one figure in 2 rows, and then save the entire figure.
import matplotlib.pyplot as plt
fig, ([ax1, ax2, ax3], [ax4, ax5, ax6]) = plt.subplots(3, 2, shareX = True, shareY = True)
fig.subplots_adjust(hspace = 0, wspace = 0)
scatter([1,2,3], [4,5,6])
scatter([7,8,9], [1,2,3])
scatter([4,5,6], [7,8,9])
scatter([3,2,1], [4,5,6])
scatter([9,8,7], [1,2,3])
scatter([6,5,4], [7,8,9])
plt.show
plt.savefig('test')round brackets around [ax1, ax2, ax3] & [ax4, ax5, ax6] shareX -> sharex & shareY -> sharey
ax1., ax2., … missing before scatter(…)
round bracktes after plt.show missing
must be ‘test.pdf’, not ‘test’
5.3 Basics - list comprehension [05:01]
list comprehension
Häufig verwenden wir eine Schleife, um eine Operation auf die einzelnen Elemente einer Liste anzuwenden, und das Ergebni in eine neue Schleife zu schreiben. Für den Fall einer kurzen Schleife gibt es eine Kurzschreibweise, welche den Code knackig verkürzt.
It is a mathematical term, when a set is comprehended, i.e., constructed or defined, which has a similar structure to the list comprehension.
✓True
✗False
✓True
✗False
[['Chantal', 'Schmidt'], ['Thandiwe', 'Nkosi'], ['Joe', 'McIntyre']] to ['Chantal Schmidt'], ['Thandiwe Nkosi'], ['Joe McIntyre'].
float() simply converts the numpy output object into a sensible float number output.
list comprehension must be in square brackets
round(float(np.sin(n)),2) and n need to be the other way round
range must be in round brackets
5.4 Statistics - mean, median, stdev, moving average [22:38]
mean, median, standard deviation, moving average, list comprehension
Zum Einstieg in statistische Methoden gibt es einen ersten Blick auf Mittelwert, Median, Standardabweichung und gleitenden Durchschnitt (moving average).
max_points = 10000
batch_size = 20
random_numbers = np.random.rand(max_points)
calc_mean = np.mean(random_numbers)
calc_median = np.median(random_numbers)
calc_std = np.std(random_numbers)
moving_mean = [np.mean(random_numbers[i - batch_size:i]) for i in range(len(random_numbers))]
plt.plot(random_numbers)
plt.plot(moving_mean, c='orange', linestyle='dashed')
plt.axhline(calc_mean, c='grey', linestyle='dashed')
plt.axhline(calc_median, c='darkgrey', linestyle='dashed')
plt.axhline(calc_mean - calc_std, c='y', linestyle='dashed')
plt.axhline(calc_mean + calc_std, c='y', linestyle='dashed')
plt.xlabel('number of points')
plt.xlim([0,max_points])
plt.show()/Users/dominik/anaconda3/lib/python3.11/site-packages/numpy/core/fromnumeric.py:3504: RuntimeWarning:
Mean of empty slice.
/Users/dominik/anaconda3/lib/python3.11/site-packages/numpy/core/_methods.py:129: RuntimeWarning:
invalid value encountered in scalar divide
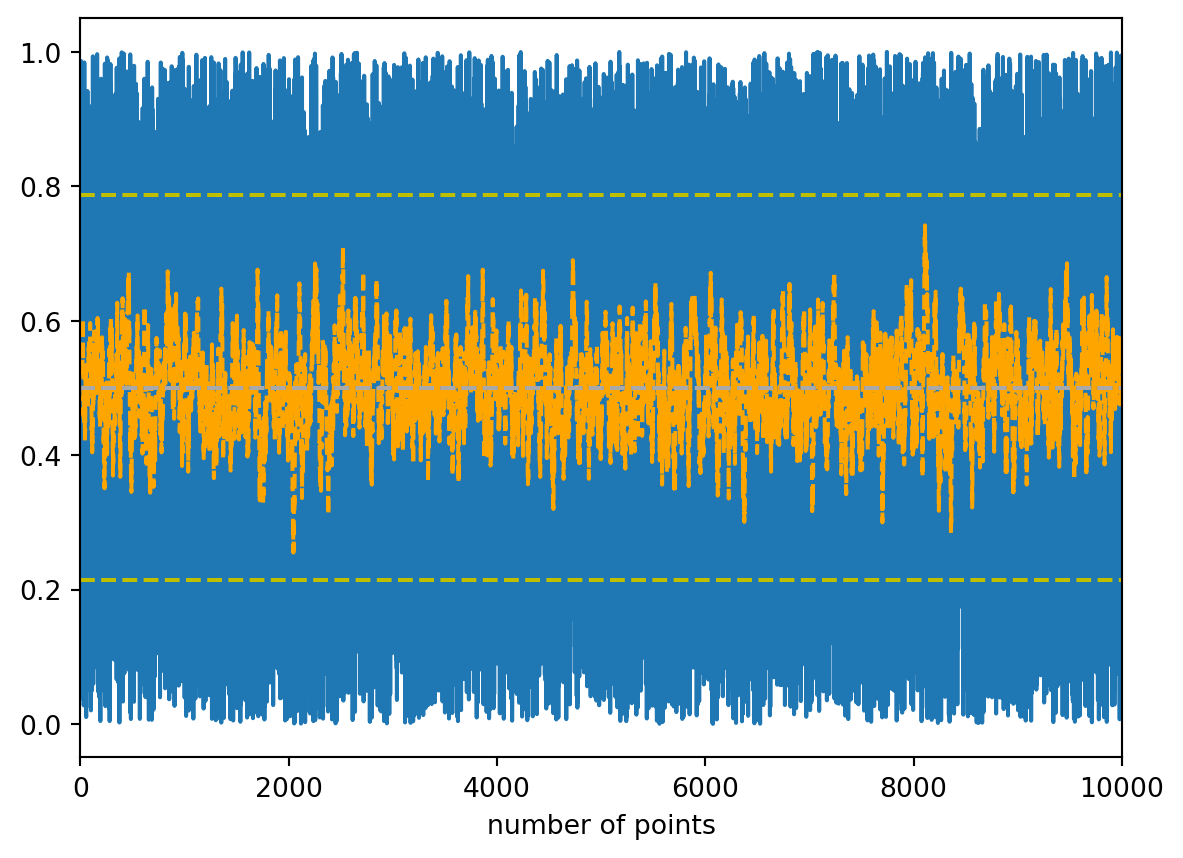
It is mean of the previous x (with x being any number) values at each point, time intervall, or the like.
✗True
✓False
✗True
✓False
Simply replace np.mean with np.median - and rename the respectvive variables accordingly.
max_points = 10000
batch_size = 20
random_numbers = np.random.rand(max_points)
calc_mean = np.mean(random_numbers)
calc_median = np.median(random_numbers)
calc_std = np.std(random_numbers)
moving_median = [np.median(random_numbers[i - batch_size:i]) for i in range(len(random_numbers))]
plt.plot(random_numbers)
plt.plot(moving_median, c='orange', linestyle='dashed')
plt.axhline(calc_mean, c='grey', linestyle='dashed')
plt.axhline(calc_median, c='darkgrey', linestyle='dashed')
plt.xlabel('number of points')
plt.xlim([0,max_points])
plt.show()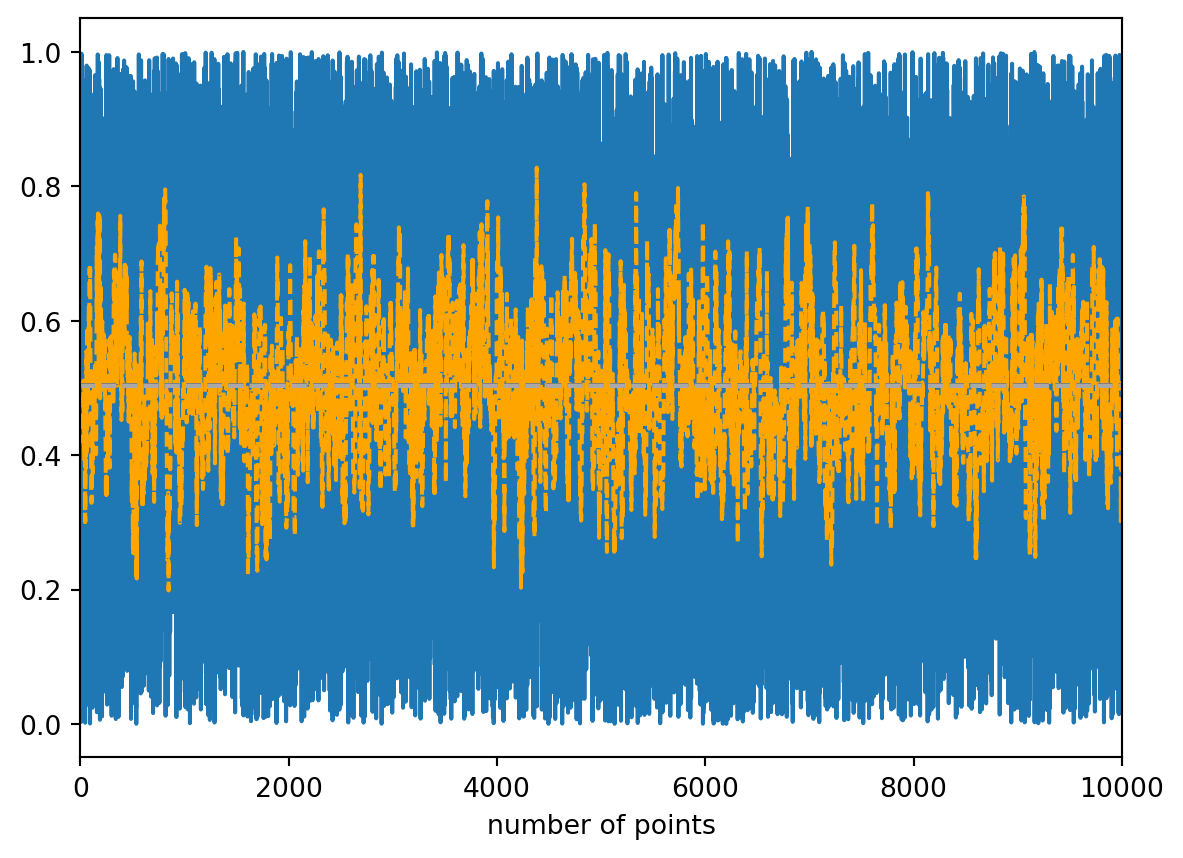
'10' must be 10, i.e., a number not a string
must be np.mean(...), not mean(...)
must be i-batch_size not i+batch_size, in the first case the mean of the next i data is taken, not the previous i data, as is correct and done in the second case, and which would then also cause incorrect results with respect to range(batch_size, len(data))
data must be i
5.5 Statistics - example of a moving average [07:18]
moving average, stocks
Als ein bekanntes Beispiel mit einfachem online-Zugriff schauen wir uns den moving average von Aktien-Preisen an.
import yfinance as yf
import pandas as pd
import matplotlib.pyplot as plt
from datetime import datetime
batch_days = 365
today = datetime.today().strftime('%Y-%m-%d')
ticker = 'AAPL'
stock_data = yf.download(ticker, start='2020-01-01', end=today)
plot_data = [stock_data['Close'][i-batch_days:i].mean() for i in range(len(stock_data['Close']))]
plt.plot(stock_data.index, stock_data['Close'])
plt.plot(stock_data.index, plot_data)
plt.show()YF.download() has changed argument auto_adjust default to True[*********************100%***********************] 1 of 1 completed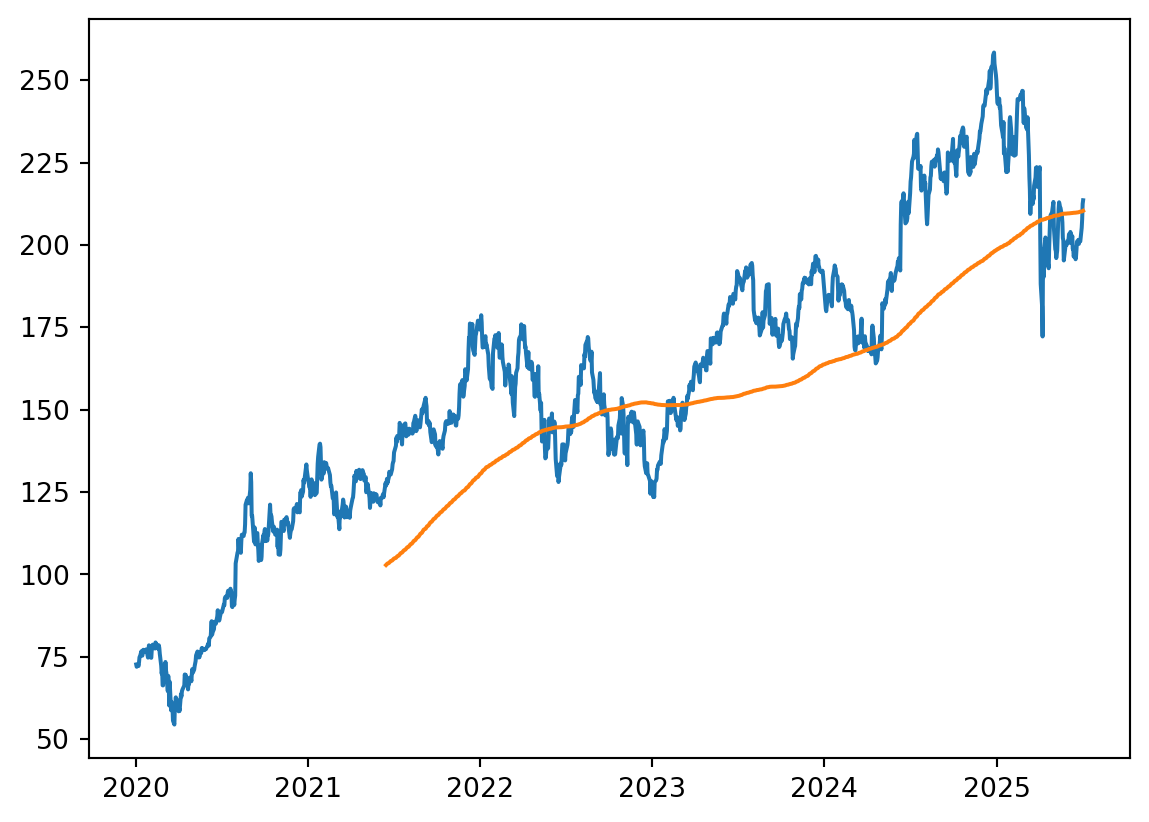
Berkeley Earth is a good resource for that matter. The data can be donwloaded or directly used as convenient text-files
✓True
✗False
✓True
✗False
Copy the content of the first 2 cells to your notebook, run them, and start with the df defined at the end of these to solve this practise. This df contains a ready to use global, historic temperature dataset.
The solution video further below explains the entire code in detail.
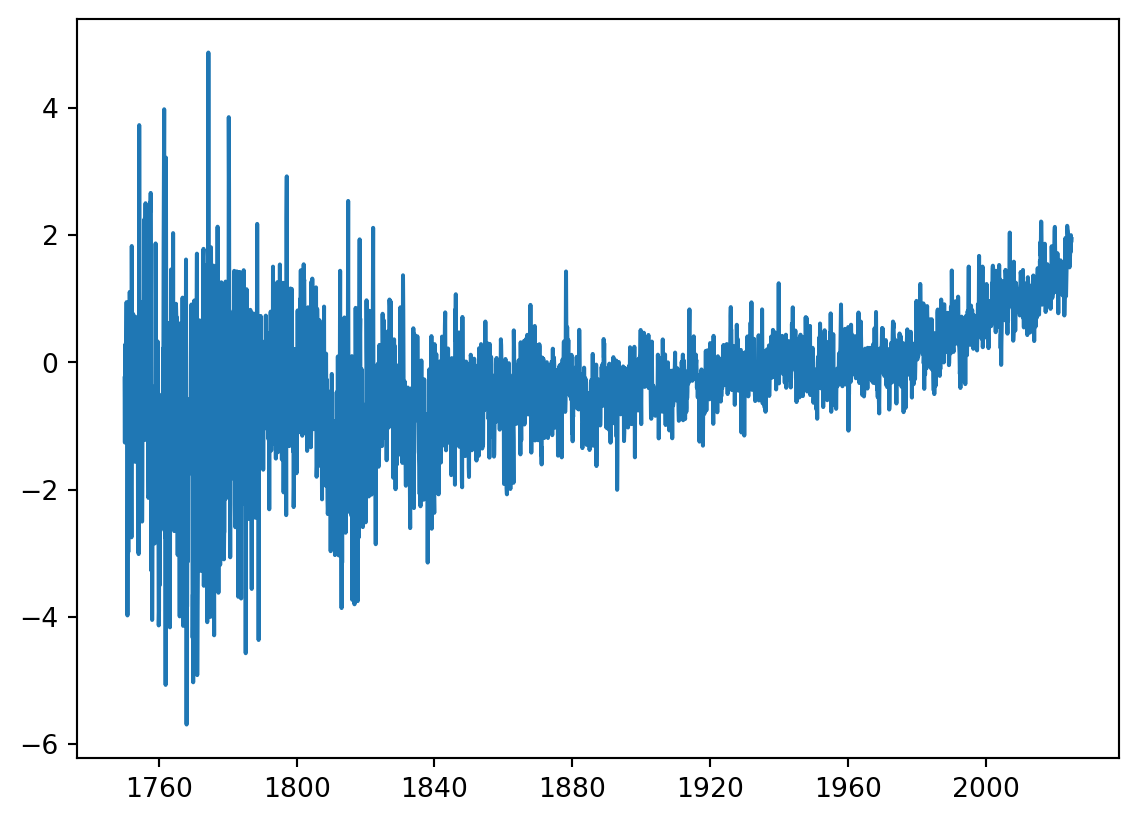
Donwloading and plotting the temperature data only.
url = "https://berkeley-earth-temperature.s3.us-west-1.amazonaws.com/Global/Complete_TAVG_complete.txt"
df = pd.read_csv(url, comment="%", sep='\s+'
,names=['Year', 'Month', 'Anomaly']
,usecols=[0, 1, 2])
df["date"] = pd.to_datetime(dict(year=df['Year'], month=df['Month'], day=15))
df = df[['date', 'Anomaly']]
df.head()| date | Anomaly | |
|---|---|---|
| 0 | 1750-01-15 | -0.252 |
| 1 | 1750-02-15 | -1.261 |
| 2 | 1750-03-15 | 0.225 |
| 3 | 1750-04-15 | 0.288 |
| 4 | 1750-05-15 | -0.970 |
Adding the moving temperature average.
The slider is not evaluated in this online book, i.e., there is no change on the plot when sliding the slider up or down.
def global_T_mov_av(batch_size):
data = df['Anomaly']
data2 = [np.mean(data[i-batch_size:i]) for i in range(batch_size, len(data))]
plt.plot(df['date'], data)
plt.plot(df['date'][batch_size:], data2)
plt.xlabel('time')
plt.ylabel('global temperature anomaly (ºC)')
return plt.show()
interact(global_T_mov_av, batch_size = (1, 100, 1))<function __main__.global_T_mov_av(batch_size)>Storing the downloaded and manipulated data locally in your file system und the specified path, if desired.
coming soon
pip install mag4
We usually use interact(sc_plot, xEl=('Si', elements), yEl=elements). I modified this slightly using widgets (a package I import above together with interact), so an initial value for the dropdown menues can be preselected. There is no error in this modified, new part.
import matplotlib.pyplot as plt needs to be imported
must be data.columns.tolist()[27:], not columns.tolist()[27:]
this is a tricky one: as the data are loaded into data, not df, it must be plt.scatter(df[xEl]/10000, df[yEl]/10000), not plt.scatter(data[xEl]/10000, data[yEl]/10000)
everything in the def needs to be indented
plt.show() should be return plt.show() (although it works both ways)